Getting Data from NCBI¶
learning-objectives
- Know what the NCBI Sequence Read Archive is
- Know what information about an experiment is available on the SRA
- Know what metadata is and how it is important to an experiment
- Know what files are available from NCBI and how they must be converted
- Be able to download NCBI SRA data and convert it to fastq format
NCBI Sequence Read Archive¶
The NCBI Sequence Read Archive (SRA) is a repository of DNA and RNA sequence. When researchers sequence DNA or RNA, and then publish their results, they will usually deposit a copy of of that data in the SRA. This makes it possible for other researchers to use that data and reproduce the findings of other researchers.
Tip
While the SRA contains information from genome sequencing (DNA) it also contains information from RNA-Seq experiments (RNA). However, you will not find RNA sequences (e.g. ‘AUGC…’) on the SRA. In an RNA-Seq experiment, the RNA sequence is converted to DNA using an enzyme called reverse-transcriptase. This is done because RNA is very unstable; converting it to DNA makes it easier to work with.
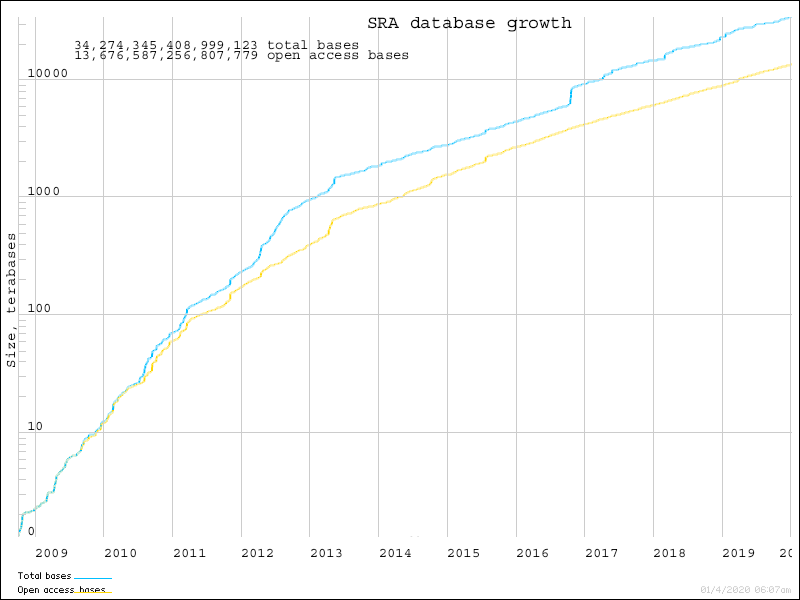
At the time of this tutorial, the SRA had more than 34 quadrillion bases of sequence!
This is a lot of data, so special tools will be needed to search and download the specific data you are looking for.
Accessing an SRA Bioproject¶
From the SRA website, you can search for data using any search term (e.g. organism type, disease, etc.). In the Leptin paper they tell us where on NCBI we can find their data.
Exercise: searching on SRA¶
- Go to the SRA website and search for the accession mentioned in the Leptin paper: PRJNA353374
You will see a description of the experiment and get a list of 12 samples related to this experiment.
- To see an experiment abstract you can go directly to the Bioproject ( which should be the link at the top of the page)
From this page we see some information describing the experiment. In the Project Data section of the page, we see there are 12 SRA experiments registered.
- Click on the SRA Experiments link (SRA experiments) to get a list of the experiments.
- Click on the first link Regular Diet Control 1 (Regular Diet Control 1)
There are a number of important things we can learn from this page:
- These data were prepared on an Illumina HiSeq 2500 sequencing machine
- This is an RNA-Seq experiment
- The Layout is SINGLE (meaning only one strand of double-stranded sequence was generated)
These and other information are the experiment’s metadata - detailed information about the experiment that we will need for our analysis.
- We can see the metadata for all the experiment using the NCBI run selector; Under Study click the All runs link (All runs).
The All runs data gives us detailed information about the experiment. At the bottom of the page is a table with the ID (run) for each of the samples. In the Sample Name column we see how each sample is categorized. Samples end in a number which is the replicate (we take multiple samples called replicates from mice different mice undergoing the same treatment). In the treatment column we see additional details on the mice that yielded the samples.
Downloading and converting data from SRA¶
Using the Jupyter Notebook, you will be downloading data from SRA. There is no “download” button on SRA. Instead, there is special software called SRA tools that allows you to download data from SRA. Data download from the SRA will be compressed and in the SRA format with the “.sra” file extension. Most bioinformatics tools cannot read these files, so we will also use sra tools to convert the data into the fastq format. Fastq is the most common format for high-throughput DNA sequencing (the type of sequencing generally used for genome sequencing and also RNA-Seq). Most tools are compatible with this format.
Access the JupyterLab Lesson on CyVerse and complete Notebook 0: Import file from NCBI Sequence Read Archive
Notebook Preview
This is a preview of the notebook in this lesson. Go back to JupyterLab Lesson on CyVerse to launch and use the interactive notebook.
Questions¶
Question
- What is the SRA?
- What is metadata, and what are some things SRA metadata might tell you about an experiment?
- What format do SRA files use to store sequence data?
- What format do SRA files need to be converted to?
- How can SRA files be converted?